4. Compute chokepoints¶
Given a genome-scale model in SBML format, findCPcli computes chokepoints, dead-end metabolites, essential reactions, and essential genes, and saves the results in a spreadsheet. findCPcli can be run as follows, where model.xml is the file with the SBML model and results.xls is the file where the results will be saved.
$ findCPcli -i model.xml -o results.xls
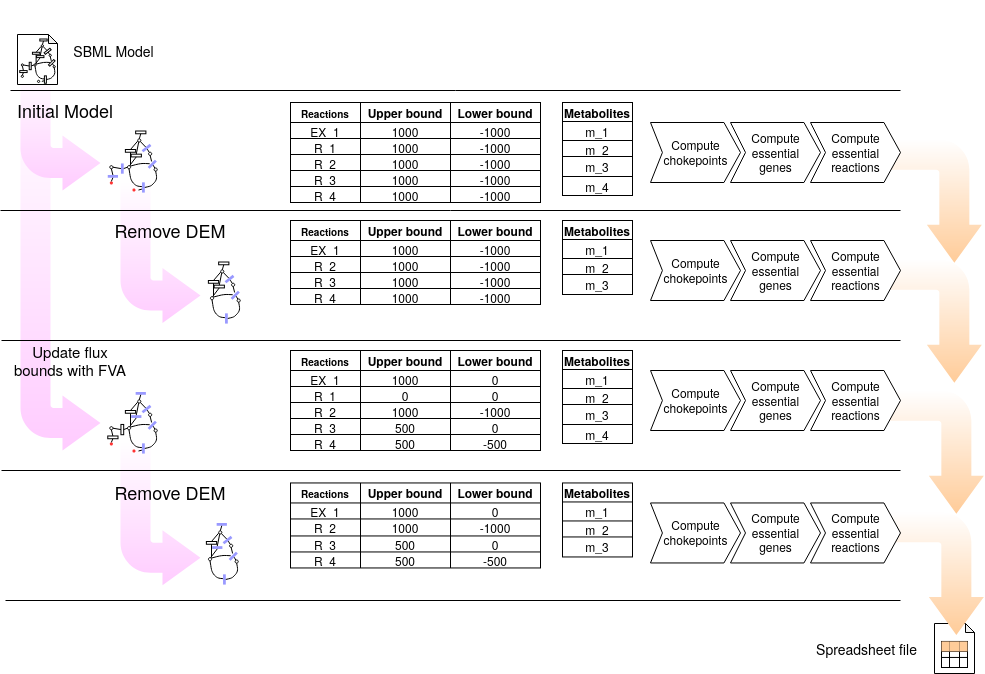
The following figure shows the pipeline of the chokepoint computation process. For a given SBML file, computations are performed on 4 models: i) model in the SBML file; ii) model without DEM; iii) model refined with FVA, i.e. with flux bounds updated according to FVA; and iv) model refined with FVA and without DEM.
4.1. Spreadsheet data¶
The previous command produces a spreadsheet file containing the following sheets:
model_info: General model information.reactions: List of reactions of the modelmetabolites: List of metabolites of the modelgenes: List of genes of the modelreactions_FVA: Upper and lower flux bound of each reaction obtained with Flux Variability Analysis.metabolites_FVA: Upper and lower flux bound of each reaction obtained with Flux Variability Analysis grouped by metabolite.reversible_reactions: List of reversible reactions of the model before and after FVA refinement.chokepoints: Chokepoint reactions and the metabolite/s they produce/consume. Chokepoints are computed in 4 different models:Input model in the SBML file.
Model without DEM.
Model refined with FVA.
Model refined with FVA and without DEM.
dead-end: Dead-end metabolites before and after FVA refinement.essential genes: List of essential genes of the model. Essential genes are computed in the 4 previously listed models.essential reactions: List of essential reactions of the model. Essential reactions are computed in the 4 previously listed models.comparison: Comparison of chokepoint, essential reactions and essential gene reactions in the 4 previously listed models.summary: Comparison the size of the previous sets and their intersections.